exatlas-help file.
Two main types of data files are gene expression
profiles and gene sets, which can be uploaded manually or retrieved from GEO database.
Tools for comparison of two or more data sets are shown as yellow boxes.
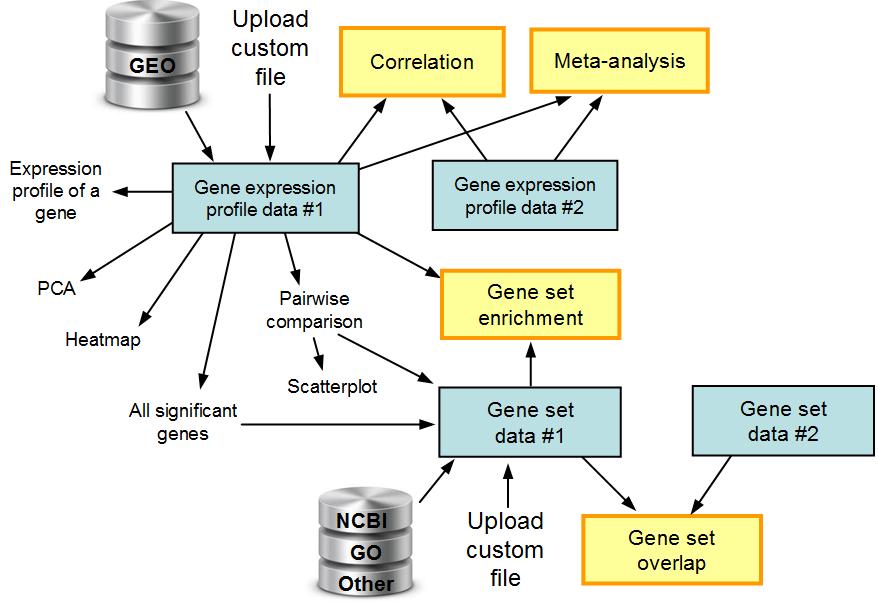
Fig. 1. The workflow in ExAtlas.
Gene expression data can be uploaded manually as tab-delimited text files, which are normalized
as transcripts per million (TPM) during the upload. Preloaded public data sets
improve the functionality of the software because they save time on data
search, retrieval, and preprocessing.
Users can store their data files indefinitely in their account and keep using them for
additional analyses. The file management system allows users to download, delete, or edit their
data files.
ANOVA is used in ExAtlas to evaluate the variability of log-transformed gene expression
values between replications. First, one-way ANOVA is applied to each individual gene (or probe in the array),
and then the error variance (also known as mean square error) is adjusted to the results
obtained for other genes. Error variance may appear very low by pure chance for
some genes, especially if the number of replications is small; and this factor may increase
the number of false positives. Error models used in ExAtlas are designed to get a better estimate for
the true error variance than the error variance estimated from expression of a single gene
(i.e., empirical error variance). Adjustment is done by taking the maximum of the empirical error variance
for a single gene and the average error variance for
multiple genes that are expressed at approximately the same level as the given gene.
The average errov variance is estimated as follows.
All genes are sorted based on the average log-expression in all samples,
and then the average error variance is estimated in a sliding window of 500 genes.
The top 1% of genes with highest empirical error variances are not counted in this calculation
to avoid potential bias due to outliers and highly variable genes.
Low-expressed genes generally have higher error variances than high-expressed
genes. However, their variation could be underestimated due to data processing (e.g., using
cutoff thresholds, rounding, or pseudo-values). Thus in ExAtlas, the average error
variance is forced to increase monotonically with decreasing expression of genes below
the median. According to the default error model, adjusted error variance equals the
maximum of the empirical error variance and the average error variance. Other error
models are available when users select the option "Run ANOVA again." These include
using empirical error variance without adjustment, or average error variance, or Bayesian
correction of error variance, or the maximum between empirical error variance
and its Bayesian correction.
With ExAtlas, users can run ANOVA-like analysis even for data sets with no replications
(this option is usually not available in other software). In this case, the error variance
is estimated based on the half-normal probability plot method. We assume that at least
a half of degrees of freedom in a set of gene expression values
represent random effects. Thus, the standard deviation, σ, of random effects can be
approximated by the median of positive deviation (i.e., absolute value of deviation) from the
mean divided by 0.675 (inverse half-normal cumulative distribution for p=0.5). The error variance in ANOVA is then set to σ2.
This method is applied to each set of the 500 genes in a sliding window that is shifted
across the set of all genes sorted by their average log-expression. This error variance is
then used for evaluating the significance of gene expression change in individual genes.
Specificity of gene expression in a specific tissue or cell type is evaluated by a z-value
that compares expression in a given tissue with the average expression in other tissues
that are not correlated with the tested tissue. Tissues are considered correlated if the
log-transformed multidimensional distance of their log-transformed expression profiles
is less than 1/3 of the maximum distance between
tissues measured by PCA. Specificity of a gene is considered significant if z > 2.
Heatmap and PCA are used to explore gene expression profiles of all tissues or
cell types in a data set. These methods can be applied to a subset of genes that satify
criteria of significance (thresholds of FDR and fold change). By
default, both the heatmap and PCA are applied to averages of log-transformed gene
expression in each tissue, but there is an option to use individual replications for analysis.
PCA determines major correlations among gene expression profiles in a data set. Users can
check the box "PC gene clustering" to identify genes that are correlated positively and
negatively with each principal component. In the heatmap, both columns and rows
are sorted by hierarchical clustering. However, if the number of genes is >2000, then
rows (genes) are clustered by PCA: first by positive and negative values of each PC
that exceed 1/4 of the maximum value, then by combinations of values in each
pair PCs (both >1/4 of the maximum value and the ratio between PCs < threefold).
Residual genes that matched neither to individual PCs nor to their pairs are placed at the
end of the heatmap and sorted by PC1.
Correlation between gene expression profiles can be identified in ExAtlas in order to
detect functional similarity
between cell types or tissues. Thus, it helps in functional annotation of transcriptome
changes. For example, the function of various transcription factors (TFs) can be inferred
from the expression change of downstream genes after induction (or repression) of these
TFs in cultured pluripotent cells (embryonic stem cells ESC or induced pluripotent stem
cells iPSC). Such experiments has been done with several humdreds of mouse and human TFs,
and the results are included as public resources in ExAtlas.
Genes that do not show significant change of expression may add noise to
correlation analyses. Thus, it is reasonable to analyze only those genes with significant
change of expression in both data sets.
If the correlation between gene expression profiles is positive and significant,
then we expect that the number of genes upregulated in both profiles or
downregulated in both profiles (called "coregulated" genes) is larger than expected
in pairs of noncorrelated gene expression profiles. To identify coregulated genes and
evaluate their statistical significance, users select option "Identify coregulated genes"
before doing correlation analysis. Statistical significance is evaluated by EPFP, which
is similar to FDR but is not estimated from p-values. The algorithm for finding coregulated
genes is based on the analysis of data points in the positive quadrant (i.e., x > 0 and
y > 0) for upregulated genes, and negative quadrant (i.e., x < 0 and y < 0) for downregulated
genes, where x and y are expression values of a gene (i.e., log ratio vs. median
or other baseline) in samples compared from data sets #1 and #2, respectively. Because
the analysis of upregulated and downregulated genes is identical except for the direction
of expression change, the algorithm is described here only for the upregulated genes.
First, gene expression change values are replaced by their rank order in the positive
quadrant. If the null hypothesis holds (i.e., no correlation), then genes are expected
to have a uniform random distribution (Fig. 2B). However, if gene expression profiles
are positively correlated, then more genes (points) can be found near the upper right
corner of the plot (Fig. 2B). To estimate EPFP(rx,ry) for a gene with rank orders rx
and ry in the first and second expression profile, respectively, the density of dots/genes
is estimated in a dark-shaded rectangle in Fig. 2B, and compared to the density of dots
in two adjacent light-shaded rectangles. The EPFP equals the density of dots in the
light-shaded area divided by the density of dots in the dark-shaded area. Because we
have two light-shaded rectangles, EPFP is estimated twice, and then the
largest value is selected (to be conservative). Because EPFP may not monotonically
decrease with the increasing rank of genes, it is forced to decrease monotonically. In
particular, if EPFP(rx1,ry1) > EPFP(rx,ry), rx1 > rx, and ry1 > ry, then EPFP(rx1,ry1)
is reassigned to EPFP(rx,ry). All genes with EPFP below a selected threshold are considered
coregulated (Fig. 2C). To identify oppositely coregulated genes (i.e., upregulated
genes in file #1 that are downregulated in file #2), the direction of change in file 2 is set
to "Reversed." The gene expression change in file 2 is thus inverted (i.e., multiplied
by -1), and oppositely coregulated genes can be identified in the same way as shown
in Fig. 2.
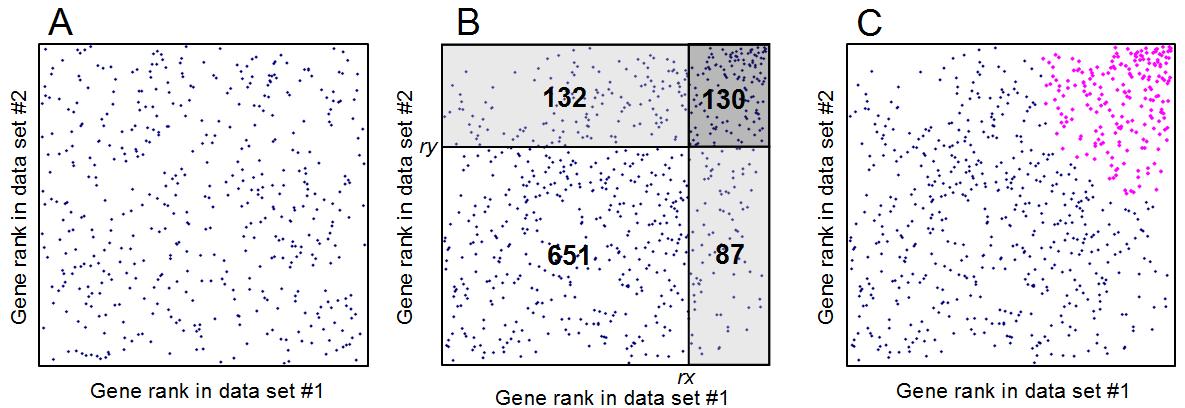
Fig. 2. Estimating the expected proportion of false positives (EPFP) for positively coregulated
genes: (A) scatterplot of gene expression ranks in the positive quadrant if there is no correlation.
(B) The same plot if gene expression profiles are correlated; numbers indicate gene counts in each
area. The density of dots/genes in the dark-shaded rectangle is compared to the density of dots in
two light-shaded rectangles used as baseline, generating two estimates of EPFP for a gene at the lower
left corner of the dark-shaded rectangle (with expression rank orders rx = 651 + 132 = 783 and
ry = 651 + 87 = 738): EPFP1 = 0.281 and EPFP2 = 0.237. The greater value of these is then selected:
EPFP = 0.281. (C) All coregulated genes with EPFP ≤ 0.3 are highlighted in magenta.
Gene set overlap and gene set enrichment analyses are used to evaluate if specific gene sets
(such as GO or KEGG) are overrepresented among upregulated and/or downregulated
genes. For example, if a set of upregulated genes, S1, is identified using threshold values
of FDR and/or fold change, then we can test if this set is enriched in genes that belong to a
set of annotated genes, S2. The proportion of genes from set S2 in the genome genes
equals p = n2/N, where n2 and N are the numbers of genes in S2 and in the whole
genome, respectively. The proportion of genes from set S2 within set S1 is equal
q = m/n1, where m is the number of common genes in S1 and S2, and n1 is the number
of genes in S1. Genes that are common in sets S1 and S2 belong to the overlap of these two
sets (i.e., S1⋂S2); thus, the goal of the analysis is to check if the overlap can (or cannot) be
explained by a random draw of genes. In particular, we can say that genes from set S2 are
enriched in the set S1 if the overlap of genes in sets S1 and S2 is greater than can be
explained by random choice. Z-values are then estimateed assuming that the proportion of common genes has a hypergeometric
probability distribution.
Gene set enrichment analysis is similar to the method of gene set overlap, but it has
a greater statistical power because it does not require defining thresholds for delineating
sets of differentially expressed genes. In particular, gene set enrichment analysis can find
significant associations with functional gene sets even if there are no significantly upregulated
genes based on standard criteria (e.g., FDR ≤ 0.05 and fold change ≥ 2). Gene set
enrichment analysis uses the order of genes sorted by the change of their expression to
identify the best threshold to delineate the set of differentially expressed genes. Among
various approaches to gene set enrichment analysis, we selected Parametric Analysis of
Gene Enrichment (PAGE) because of its simplicity and reliability. The algorithm of
PAGE determines whether the mean log-expression change, xset, in genes that belong
to a functionally annotated gene set S is significantly greater than expected from the
mean and standard deviation of log-expression change in all genes
(xall and SDall, respectively).
Rank order plot (or rank-plot) is used to visualize the results of gene set enrichment
analysis (Fig. 3). Genes are sorted according to their expression change from
downregulated at the left to upregulated at the right, and then the proportion of genes
from the set of annotated genes, S, is estimated in a sliding window of 300 genes.
Fig. 3 shows an examples of enrichment of TF targets among downregulated genes in
mouse ESCs after the knockdown (KD) of transcription repressor Rest with shRNA.
Sets of target genes of individual TFs include all genes with TF binding
sites in their promoters and enhancers, based on published ChIP-seq experiments.
When Rest is repressed, its target gens become upregulated, showing a peak
on the right in Fig. 3.
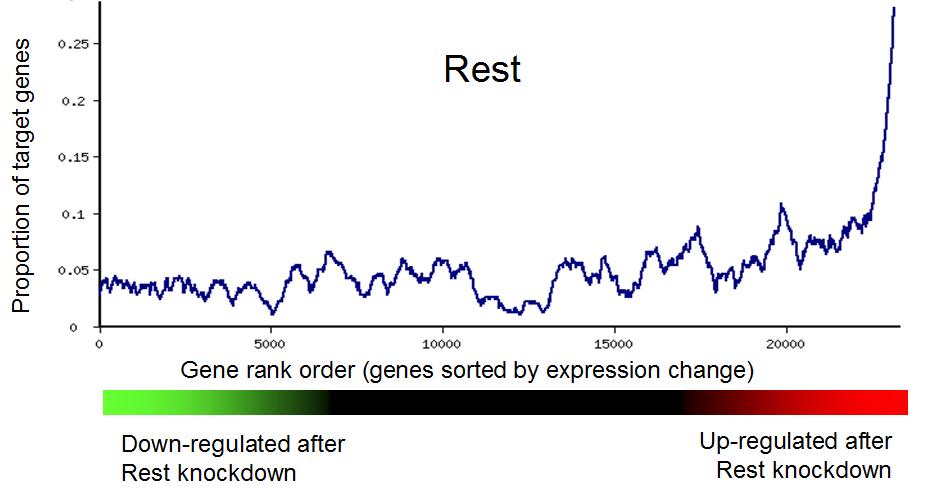
Fig. 3. Gene set enrichment analysis is used to generate a rank plot of
the proportion of target genes of transcription factor Rest among genes
that were downregulated and upregulated after the knockdown of Rest.
If gene set enrichment is statistically significant (P ≤ 0.05) among upregulated genes,
then some genes in the annotated gene-set are more abundant among
upregulated genes than expected by random. However, PAGE does not provide information
on the EPFP in these genes. Most studies of downstream effects of TFs assume
that the set of regulated target genes is the intersection of two subsets: a subset of all target
genes and another of genes that changed their expression following manipulation of a TF.
However, the intersection may include a large number of false positive genes
that change their expression not because of TF binding to promoters or enhancers but
from other causes (e.g., second-order effects, compensation, etc.). Thus, a smaller
subset of genes within the intersection needs to be delineated to bring EPFP below an
acceptable threshold (Fig. 4). To identify enriched genes and evaluate their statistical
significance by EPFP, users check a box "Identify associated genes" before doing gene set
enrichment analysis. EPFP is calculated sequentially for different thresholds of gene
expression change as a ratio of the proportion of targets among "control" genes (i.e.,
not affected by TF manipulation) to the proportion of targets among genes that
responded to TF manipulation. The smallest fold change threshold that yields EPFP
below the accepted level (e.g.EPFP ≤ 0.3), is then used to delineate the set of regulated targets
of TFs.
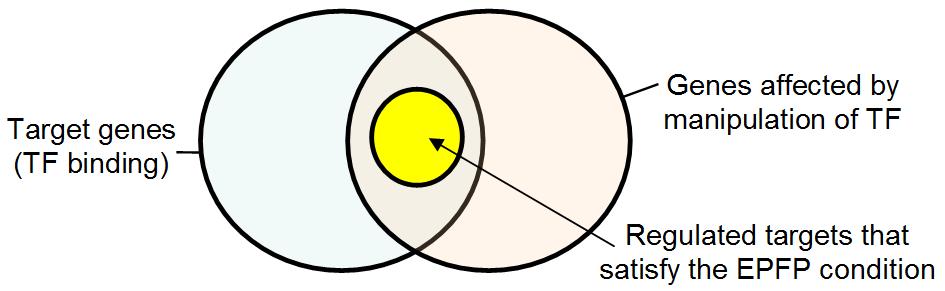
Fig. 4. Regulated targets of TFs represent a subset of genes within the set of common genes that
are both TF tragets and upregulated (or dopwnregulated) after TF induction (or repression.
Methods of gene set analyses described earlier assume that all genes in a set are
equally weighted. However, experimental and mining methods used for generating
gene sets often provide additional quantitative information for individual genes. Genes
in a set may differ by z-score, p-value, expression change, number of sequence reads in
ChIP-seq peaks, or in HITS-CLIP assay, and other criteria that may appear useful for
the analysis of gene set enrichment. Thus, an option to handle gene sets with additional
quantitative attributes was added to ExAtlas. If the box "Use attributes" is checked,
then genes with low scores are transiently excluded from the analysis for varying score
thresholds until the highest statistical significance (z-value) is achieved. This approach is
applied both to gene set enrichment (PAGE) and to gene set overlap (hypergeometric
test).
The goal of meta-analysis is to use information from multiple independent studies to
increase the power of statistical inference and reduce the number of false positives.
ExAtlas applies four frequently used methods of meta-analysis: Fisher s, z-score, fixed
effect, and random effect to evaluate the difference in gene expression between two
Figure 4.6 Gene set enrichment scores (PAGE) for Gene Ontology (GO) annotated sets of genes
among upregulated genes in mouse ESCs after induction of 137 individual transcription factors
(TFs). Columns, induced TFs; rows, GO annotations.
tissues or cell types. The first three methods assume that all studies implement exactly
the same methodology (e.g., same cell lines, equipment, and reagents). In practice,
however, methods often do not match between experiments, so the random effect
seems to be most relevant method. Fisher s method combines log-transformed p-values
from m studies and generates a chi-square statistic with 2*m degrees of freedom. The
z-score method combines z-scores (i.e., the ratio of mean effect to its SD) of different
studies with weights equal to square root of sample size. The fixed effect method estimates
a weighted sum of effects (i.e., log ratio of gene expression change), where
weights are equal to 1/σ2 with σ2 as the error variance. The random effect method takes
into account the variance of heterogeneity between studies, T2, which is added to the
variance of individual effects.
Meta-analysis in ExAtlas starts by selecting gene expression data to be combined. Pairs
of tissues (or cell types) are compiled from different data files or the same file. It is even
possible to compile gene expression data from different species, and in that case, gene
orthologs are compared. A list of all expression profile pairs used in meta-analysis can
be saved for future retrieval and modification. The analysis is done using all four methods,
and a table with the number of significantly overexpressed and underexpressed genes for
each method is displayed. Differentially expressed genes can be saved as gene sets for
functional annotation.

